- Code: Select all
//Adaptation of Albert Jan Wonnink's GAViewer code for the S^3 model of the singlet correlations
function getRandomLambda()
{
if( rand()>0.5) {return 1;} else {return -1;}
}
function getRandomUnitVector() //unit vectors uniformly distributed over S^2
//http://mathworld.wolfram.com/SpherePointPicking.html
{
v=randGaussStd()*e1+randGaussStd()*e2+ randGaussStd()*e3; //three-dimensional vectors
return normalize(v);
}
batch test()
{
set_window_title("Test of 3D S^3 GA Model for the 2-particle singlet correlations");
default_model(p3ga); //choice of the model in GAViewer
N=10000; //number of iterations (trials)
I=e1^e2^e3; //the fundamental trivector of GA
ss=0;
t=0;
u=0;
for(nn=0;nn<N;nn=nn+1) //perform the EPR-Bohm experiment N times
{
a=getRandomUnitVector();
Da=I a; //detector bivector of Alice as in Eq.(30)
b=getRandomUnitVector();
Db=I b; //detector bivector of Bob as in Eq.(31)
s1=getRandomUnitVector();
s2=s1; //conservation of zero spin as in Eq.(43)
Ls1=I s1; //bivector representing spin of particle 1
Ls2=I s2; //bivector representing spin of particle 2
lambda=getRandomLambda(); //lambda is a fair coin, giving the +1 or -1 choice
A=(-Da*lambda*Ls1); //Alice's measurement function as in Eq.(30)
B=(lambda*Ls2*Db); //Bob's measurement function as in Eq.(31)
//Note that the limits on A and B are not required because of the conservation of
//spin, s2=s1, is imposed which reduces the product -Ls1*Ls2 in A*B to unity.
q=0;
if(lambda==1) {q=A B;} else {q=B A;} //shuffles the alternative orientations of S^3
ss=ss+q;
phi_a=atan2(scalar(Da/(e3^e1)), scalar(Da/(e2^e3))); //gets azimuthal angle for a
phi_b=atan2(scalar(Db/(e2^e3)), scalar(Db/(e3^e1))); //gets -azimuthal angle for b
neg_adotb=-(a.b);
print(neg_adotb, "f"); //outputs -a.b event by event
if(phi_a*phi_b>0) {eta_ab=acos(a.b)*180/pi;} else {eta_ab=-acos(a.b)*180/pi+360;}
print(eta_ab, "f"); //output the angles eta_ab event by event
print(correlation=scalar(q), "f"); //output the correlations; cf. Ref.[22] & Fig. 6
t=t+A;
u=u+B;
}
mean=ss/N;
print(mean, "f"); //shows the vanishing of the non-scalar part
aveA=t/N;
print(aveA, "f"); //verifies that individual average < A > = 0
aveB=u/N;
print(aveB, "f"); //verifies that individual average < B > = 0
prompt();
}
Typical output is,
- Code: Select all
neg_adotb = 0.010721
eta_ab = 269.385742
correlation = 0.010721
neg_adotb = 0.585568
eta_ab = 234.156906
correlation = 0.585568
neg_adotb = -0.176435
eta_ab = 79.837830
correlation = -0.176435
neg_adotb = 0.740447
eta_ab = 137.769531
correlation = 0.740447
neg_adotb = 0.350221
eta_ab = 110.500862
correlation = 0.350221
neg_adotb = 0.949493
eta_ab = 198.287735
correlation = 0.949493
neg_adotb = 0.313786
eta_ab = 108.287560
correlation = 0.313786
neg_adotb = -0.423052
eta_ab = 64.972549
correlation = -0.423052
neg_adotb = 0.661114
eta_ab = 228.615097
correlation = 0.661114
neg_adotb = 0.213105
eta_ab = 257.695618
correlation = 0.213105
neg_adotb = -0.421566
eta_ab = 294.933502
correlation = -0.421566
neg_adotb = -0.072056
eta_ab = 85.867928
correlation = -0.072056
neg_adotb = -0.277978
eta_ab = 286.139557
correlation = -0.277978
neg_adotb = 0.246896
eta_ab = 104.293915
correlation = 0.246896
mean = 0.001890 + 0.013989*e2^e3 + -0.004283*e3^e1 + -0.006126*e1^e2
aveA = 0.006420 + 0.007764*e2^e3 + 0.000817*e3^e1 + -0.000506*e1^e2
aveB = -0.005920 + -0.004768*e2^e3 + 0.001920*e3^e1 + 0.010372*e1^e2
And here is the plot of the data.
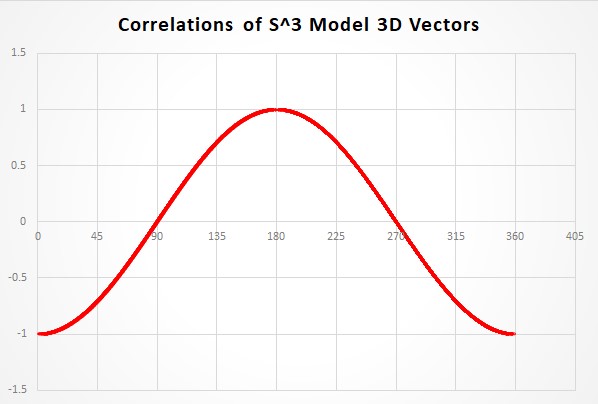
The plot of the data is very simple. The x axis is
Here is a link to the raw GAViewer data for 25K trials as a .csv file (about 500K file size). The first column is the angle and the second column is the correlation value at that angle.
EPRsims/newjoyGA3D25k2.csv
.

